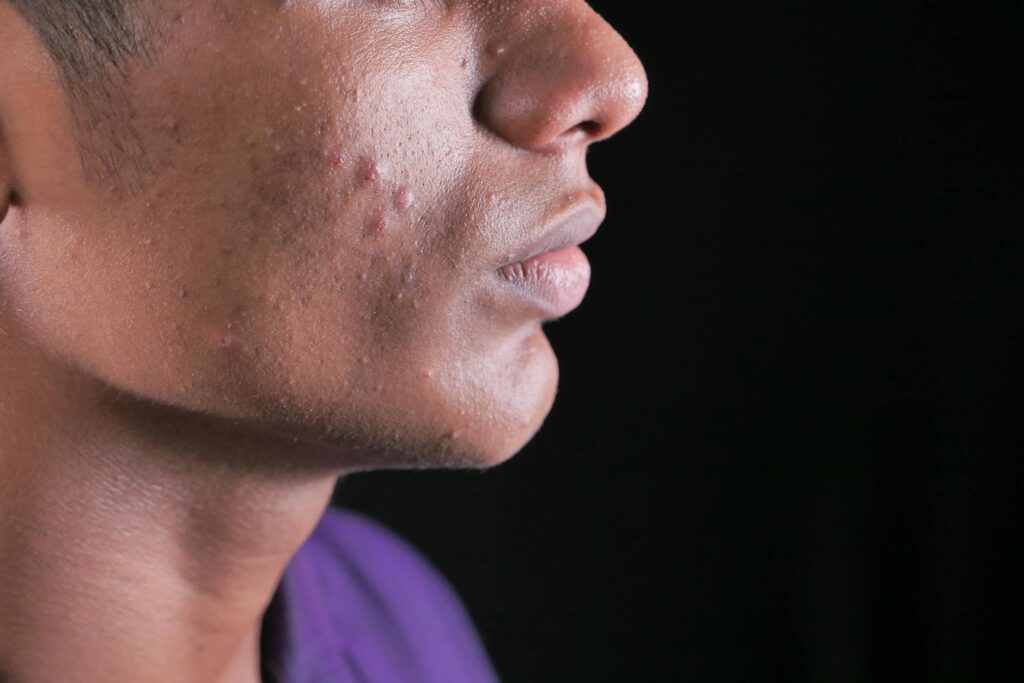
Metagenomics—the study of genetic material directly sampled from skin without culturing bacteria—reveals that acne-affected skin hosts a fundamentally different microbial landscape than healthy skin. The research shows that people with acne actually have higher species-level bacterial diversity overall, though this difference is subtle and not statistically significant (Shannon index: 0.63 for acne versus 0.44 for healthy individuals). More importantly, specific bacteria shift in abundance, virulence patterns change, and the metabolic activity of the microbiome diverges significantly from healthy skin.
These findings challenge the older narrative that simply blamed Cutibacterium acnes (formerly Propionibacterium acnes) for acne, showing instead that the entire microbial ecosystem—including unexpected players like Staphylococcus epidermidis—shapes whether skin breaks out. This article explores what large-scale metagenomic studies have uncovered about the bacterial underpinnings of acne. We’ll examine which bacteria actually dominate acne-affected skin, how specific strains matter more than the species themselves, why virulence factors matter more than bacterial abundance alone, and how the gut microbiome connects to facial breakouts. Understanding these findings helps explain why treating acne requires more nuance than simply targeting one bacterium.
Table of Contents
- How Metagenomic Sequencing Reveals Differences Between Acne and Healthy Skin
- Staphylococcus epidermidis—The Underappreciated Acne Player
- Strain-Level Patterns—Why Which Type of P. acnes Matters
- Virulence Factors and Metabolic Changes—Why Abundance Doesn’t Tell the Full Story
- Oxygen Availability and Bacterial Competition—The Missing Environmental Factor
- The Gut-Skin Axis—Systemic Connections to Acne Bacteria
- What This Means for Future Acne Treatment
- Conclusion
How Metagenomic Sequencing Reveals Differences Between Acne and Healthy Skin
Metagenomic analysis works by extracting all DNA directly from skin samples and sequencing it, bypassing the limitations of culture-based methods that can only grow a fraction of skin bacteria in the lab. When researchers applied this approach to compare acne-affected skin to healthy skin, they found substantial differences in the total number of optimized reads and unique sequences—metrics that reflect both the quantity and variety of microbial genetic material present. The acne group showed notably reduced counts in both measures compared to healthy controls, indicating a measurably different microbial ecosystem at the DNA level.
The higher diversity in acne skin (Shannon index 0.63 versus 0.44) might initially seem counterintuitive—shouldn’t acne skin have *less* diversity if one pathogenic bacterium dominates? The explanation is that while acne patients do harbor multiple bacterial species at higher relative levels, the total count of bacterial cells and genetic sequences is actually lower. This pattern suggests that acne skin hosts more species per unit of bacterial biomass, or that certain conditions favor a broader range of bacteria while simultaneously reducing overall microbial load. This finding fundamentally changes how dermatologists think about the microbiome: acne isn’t simply an overgrowth of one “bad” bacterium, but rather an altered ecological balance.

Staphylococcus epidermidis—The Underappreciated Acne Player
For decades, research focused almost exclusively on Cutibacterium acnes as the primary acne bacterium. Metagenomic studies have shifted attention to an unexpected culprit: Staphylococcus epidermidis, which was significantly elevated in acne-affected skin compared to healthy controls. S. epidermidis is normally considered a benign skin resident, and this discovery was surprising enough to reshape current understanding of acne’s microbial basis. Unlike C. acnes, which produces acne-triggering fatty acids and biofilms, S. epidermidis contributes to inflammation through different mechanisms, including immune activation and production of pathogenic factors that can worsen breakouts. The presence of elevated S.
epidermidis in acne skin suggests that it’s not simply about one dominant bacterium but about synergistic interactions between multiple species. Interestingly, metagenomic data shows that C. acnes relative abundance remained statistically similar between acne patients and healthy individuals—the difference isn’t that acne patients have significantly *more* C. acnes, but that their skin harbors a different ratio of supportive and antagonistic bacteria around it. Conversely, a phage that attacks P. acnes (Propionibacterium acnes phage) showed higher relative abundance in healthy skin, suggesting that in healthy individuals, natural viral predators help keep C. acnes in check. This reveals an underappreciated defense mechanism: a healthy microbiome includes phages that prey on acne-causing bacteria.
Strain-Level Patterns—Why Which Type of P. acnes Matters
Beyond species identification, metagenomic studies can resolve bacterial strains, revealing that specific ribotypes of P. acnes correlate strongly with acne presence. Research identified that ribotypes RT4, RT5, RT8, and RT10 were significantly more common in acne lesions (p < 0.05), while RT6 predominated in healthy skin. This strain-level distinction is crucial: not all P. acnes are equally acne-causing, and the specific genetic variants circulating in a patient's skin predict breakout severity far better than simply detecting the species’ presence. The diversity of P.
acnes strains was higher in acne patients than healthy individuals, meaning acne-prone skin hosts multiple different genetic variants of the bacterium, each potentially contributing to inflammation through slightly different mechanisms. This finding has direct implications for treatment. A broad-spectrum antibiotic might kill all P. acnes strains equally, but understanding that RT4, RT5, RT8, and RT10 are the problematic variants opens possibilities for more targeted approaches—theoretically, therapies could be designed to selectively suppress acne-prone strains while preserving RT6 and other non-pathogenic variants. Currently, no strain-specific treatments exist in mainstream dermatology, but this data forms the foundation for future precision approaches. For patients, the key takeaway is that acne’s bacterial basis is more specific than “too much C. acnes”—it’s about having the *wrong* types of bacteria and bacterial variants.

Virulence Factors and Metabolic Changes—Why Abundance Doesn’t Tell the Full Story
One of the most revealing findings from metagenomic studies is that the acne group showed enriched virulence-associated factors alongside a reduced abundance of metabolic synthesis genes. In other words, acne-affected skin hosts bacteria that are metabolically less productive but express more aggressive, damage-causing compounds. Virulence factors predominantly came from Staphylococcus species, meaning that while S. epidermidis abundance increased, its expression of pro-inflammatory and tissue-damaging factors increased even more sharply. This disconnect between bacterial count and actual harm is critical: having more of a “bad” bacterium isn’t what matters as much as having bacteria that express virulence genes.
The reduction in metabolic synthesis genes means that acne-prone skin bacteria produce fewer beneficial metabolites—compounds like short-chain fatty acids and antimicrobial peptides that normally protect skin barrier function. A healthy skin microbiome synthesizes these protective molecules continuously, but in acne skin, the metabolic capacity for this production is diminished. Recent 2025 research using metatranscriptomics (which measures actual gene expression, not just gene presence) revealed that while C. acnes dominates skin metagenomes at most sites, its contribution to actual metabolic activity is relatively modest compared to its abundance. This means the bacteria are present but not as metabolically active as their numbers suggest, further complicating the picture of how microbes cause acne.
Oxygen Availability and Bacterial Competition—The Missing Environmental Factor
A 2025 study revealed something long overlooked: oxygen availability strongly impacts C. acnes growth and its interactions with competing skin bacteria. C. acnes is slow-growing and somewhat oxygen-averse compared to faster aerobic species, but the study showed that in certain oxygen environments, C. acnes can outcompete or suppress other bacteria through mechanisms that reduce metabolic cooperation. This finding reframes acne as partly an environmental problem—not just about which bacteria are present, but about the conditions that allow certain bacteria to dominate. In low-oxygen pilosebaceous units (hair follicles), C.
acnes thrives; in well-aerated skin, more diverse aerobic bacteria can coexist, and C. acnes remains subdued. This has practical implications that dermatologists have long intuited. Topical treatments that increase skin oxygenation (certain retinoids, chemical exfoliants, and physical debridement) may work partly by shifting the oxygen landscape in follicles, making them less hospitable to C. acnes dominance. Similarly, the reason oral antibiotics help acne—beyond their direct killing effects—may involve subtle shifts in the oxygen gradient and microbial competition landscape. However, long-term antibiotic use inevitably selects for resistant bacteria, which is why understanding the ecological mechanisms (oxygen, nutrient competition, phage predation) offers a pathway toward treatments that work *with* the microbiome rather than against it.

The Gut-Skin Axis—Systemic Connections to Acne Bacteria
While skin metagenomics reveals local bacterial patterns, emerging research has identified a surprising connection: the gut microbiome influences acne risk through both microbial and metabolic pathways. A 2024 Mendelian randomization study identified eight bacterial species in the gut that were significantly associated with acne risk, plus eight metabolic abundance pathways originating from gut bacteria that linked to facial breakouts. This suggests that the foods we eat influence the gut bacteria that produce metabolites, which then systemically affect skin health and acne susceptibility.
For example, a diet favoring certain fiber types feeds gut bacteria that produce short-chain fatty acids; these metabolites circulate throughout the body and can reduce skin inflammation. The gut-skin connection explains why some patients see acne improve dramatically with dietary changes—not because the diet directly kills acne bacteria on skin, but because changing what you eat shifts your gut microbiota, which reduces inflammatory metabolites reaching the skin. Conversely, certain dietary patterns may select for gut bacteria that promote inflammation, compounding acne through a systemic pathway. This finding elevates acne from a purely topical skin problem to a whole-body microbiome issue, suggesting that truly effective long-term acne management might require addressing gut health alongside skin treatments.
What This Means for Future Acne Treatment
The metagenomic revolution in understanding acne bacteria is driving a paradigm shift in dermatology. Instead of viewing acne as simply “too much C. acnes,” the field now recognizes it as a dysbiotic state where multiple bacteria shift, virulence factors increase, protective metabolites decrease, and ecological imbalances emerge. A 2025 systematic review in the Journal of European Academy of Dermatology and Venereology confirmed that Cutibacterium and Staphylococcus play differential yet synergistic roles in acne pathogenesis, meaning that both bacteria matter, their interactions matter, and suppressing one without addressing the other may be ineffective.
This understanding is laying groundwork for next-generation acne therapies: targeted phage therapy to selectively eliminate problematic P. acnes strains, probiotics formulated with strains that restore healthy metabolite production, topical compounds that boost oxygen gradient to suppress C. acnes dominance, and dietary interventions to reshape the gut-skin axis. Current treatments (retinoids, benzoyl peroxide, antibiotics) work reasonably well partly by accident—they alter the microbiome in ways that suppress acne, but without targeting the specific mechanisms revealed by metagenomics. Future treatments can be far more precise and potentially side-effect free.
Conclusion
Metagenomics has transformed acne from a mystery into a solvable microbiome dysbiosis problem. The research clearly shows that acne-affected skin hosts a distinctive microbial ecosystem: elevated S. epidermidis, acne-prone P. acnes strains, enriched virulence factors, reduced protective metabolites, and altered bacterial interactions.
The surprise findings—that total bacterial diversity is higher in acne, that C. acnes abundance alone doesn’t predict acne, that a phage attacking C. acnes appears protective, and that the gut microbiome influences facial breakouts—all point to acne being an ecological imbalance rather than a simple infection. For patients and dermatologists, the practical takeaway is that effective acne management requires addressing the entire microbial landscape: topical treatments that shift follicle oxygen levels, dietary changes that reshape gut bacteria, lifestyle choices that reduce skin inflammation, and increasingly, targeted therapies designed from metagenomic insights. As research continues to map the specific bacterial players and their metabolic contributions, acne treatment will move from broad-spectrum approaches toward personalized, microbiome-informed strategies that restore balance rather than simply suppressing bacteria.
You Might Also Like
- Why Skin Tape-Strip Analysis Is Used in Acne Research
- Why Retinyl Retinoate Is Considered Stable for Acne Skin
- Why Phytic Acid Is a Gentle Exfoliant for Acne Skin
Browse more: Acne | Acne Scars | Adults | Back | Blackheads